Authors: Wout Dillen
Category: Paper:Long Paper
Keywords: XML, TEI, encoding, genetic criticism, digital scholarly editing
Sequentiality in Genetic Digital Scholarly Editions. Models for Encoding the Dynamics of the Writing Process
When TEI P5 version 2.0 was published in 2011, scholarly editors who are interested in the writing process of literary works gained an important instrument for encoding their genetic Digital Scholarly Editions in TEI-conformant XML. After a long process of deliberation, this version of the TEI’s encoding schema incorporated a large number of modifications proposed by the TEI MS SIG’s Workgroup on Genetic Editions that aimed to re-evaluate the existing TEI tagset in order to facilitate the encoding of genetic phenomena (TEI Consortium, 2011). The Workgroup’s ‘Encoding Model for Genetic Editions’ (2010) reveals two major points of interest in this proposal: (1) the need for the ability to encode features of the document rather than those of the text; and (2) the need for the ability to encode time, sequentiality and writing stages in those documents’ transcriptions.
The main answer to the first point of interest was the introduction of the <sourceDoc> element (as well as its <surface> children), that was allowed to exist on the same hierarchical level as the <teiHeader>, <facsimile>, and <text> elements. Since the Text Encoding Initiative has (as its name implies) historically favoured ‘text’ over ‘document’, this can be regarded as a powerful statement to the TEI community that documents are as valuable as texts in textual scholarship, and that it should be possible to transcribe them as such. As a result, this encoding model has been gratefully adopted by editors who are taking a more document-oriented approach to the transcription of their materials – like those of the Shelley Godwin Archive (S-GA) for instance (Shelley, 2013). The question remains, however, whether the use of this vocabulary is enough to classify a Digital Scholarly Edition as a ‘genetic’ edition. While the document will take up a central position in any genetic edition, the use of the ‘Genetic Editions’ document-oriented transcription model is not a distinctive feature of the genetic edition in itself.
The Workgroup’s second point of interest (the encoding of ‘time’) is much more central to genetic criticism. In ‘The Open Space of the Draft Page’, Daniel Ferrer makes a compelling argument that ‘the draft is not a text […], it is a protocol for making a text’, comparing it to a musical score that, though by itself inherently mute, can be interpreted as a set of instructions for a future performance (1998, 261). Likewise, a draft document leaves the writer with a set of instructions that help her transport the unfinished text from one writing stage to the next. The interpretation of these instructions, and of the distinction between different versions and writing stages, is one of the most important tasks of genetic criticism. This is what makes sequentiality such a key aspect of genetic editing: only by interpreting the draft materials as an interconnected sequence of writing acts can we expose the dynamics of the author’s writing process.
There are many ways of encoding this sequentiality in the transcriptions of draft materials, across varying levels of granularity. The Workgroup’s suggestion to use the <change> element to highlight distinct revision campaigns, for instance, effectively differentiates between individual versions of the same text when they are found within a single document. As Pierazzo and André’s ‘Proust Prototype’ demonstrates, this method can even be employed to sequence individual stages within a single version (2012). Going even further, projects like the CD-ROM edition of Willem Elsschot’s Achter de Schermen (2007) and the Melville Electronic Library’s TextLab software (2009-) analyze what John Bryant has called the internal ‘revision sequences’ of individual sentences (Bryant, 2008). The danger of analyzing the writing process on this small a level, however, is that the mechanics of the writing process may start to interfere with the dynamics of that writing process. From a genetic perspective, it is more important to expose the dynamic relation between the textual elements involved in a modification (e.g. ‘this is a substitution’) than the mechanical order in which that modification was made (e.g. ‘first this word was deleted, then this other word was added’). Since the exact writing sequence of such a modification is often impossible to reconstruct with any degree of certainty, consistently analyzing and sequencing all the work’s revision sites may introduce a number of hypotheses in the edition that the editor is not necessarily comfortable with committing to.
On the other side of the spectrum, analyzing larger macrogenetic processes across documents, the ‘Encoding Model for Genetic Editions’ refers to the TEI’s ‘Graphs, Networks, and Trees’ module, suggesting to encode the relations between documents as the <arc>s between <node>s in a <graph> element. Depending on the complexity of the writing process, this <graph> may result in an intricate data structure that can be used to visualize the chronology of the writing process on a highly abstract level. For writing processes that are less complex on the macrogenetic level, however, this model may be too much pain for too little gain, as a manually designed timetable could also do the trick.
The Beckett Digital Manuscript Project’s approach to encoding sequentiality into its genetic Digital Scholarly Edition of Samuel Beckett’s works tries to seek a middle ground between these two extremes: rather than analyzing the way in which individual sentences were written, the BDMP’s encoding model allows the user to discover how those sentences were changed from version to version, across different documents. By linking related semantic clusters on the sentence level across versions, this model allows for the on the fly generation of a chronological overview of all the different versions of each sentence in the corpus. As such, this model combines the ability of comparing different versions of the same work of more macrogenetically oriented approaches with the higher granularity of more microgenetically oriented approaches.
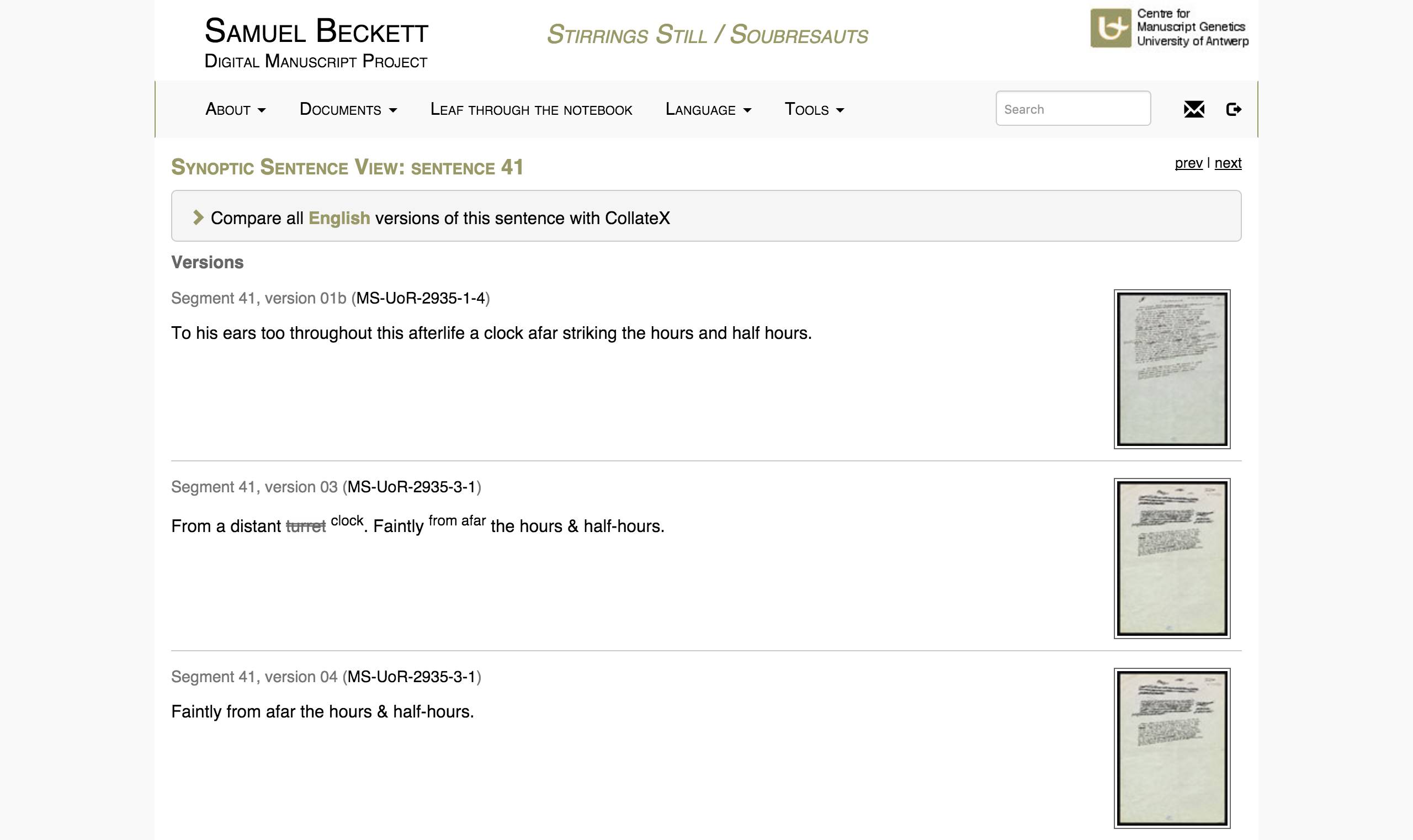
After illustrating the challenges and opportunities of these different models of encoding sequentiality in genetic editions, this paper will demonstrate how the BDMP transcribes its genetic materials in view of visualizing their sequentiality in the edition’s ‘Synoptic Sentence View’ (see ‘Figure 1’). The paper will conclude by presenting an example of how this encoding model may also be used to interpret the macrogenetic writing sequence of individual documents by means of an animated visualization of the writing process of the first draft of Beckett’s L’Innommable.
- Beckett, S. (2011). Stirrings Still / Soubresauts and Comment Dire / what is the word: a digial genetic edition (Series ‘The Beckett Digital Manuscript Project’ module 1), edited by Dirk Van Hulle and Vincent Neyt. Brussels and London, University Press Antwerp (ASP/UPA) and Bloomsbury Academic. http://www.beckettarchive.org (accessed on 12 November 2013).
- Beckett S.(2013). L’Innommable / The Unnamable: a digital genetic edition (Series ‘The Beckett Digital Manuscript Project’, module 2). Edited by Dirk Van Hulle, Shane Weller and Vincent Neyt. London and Brussels and London, Bloomsbury and University Press Antwerp (ASP/UPA) and Bloomsbury Academic. http://www.beckettarchive.org (accessed on 9 January 2014).
- Beckett S. (2015). Krapp’s Last Tape / La Dernière Bande: a digital genetic edition (Series ‘The Beckett Digital Manuscript Project’ module 3), edited by Dirk Van Hulle and Vincent Neyt. Brussels and London, University Press Antwerp (ASP/UPA) and Bloomsbury Academic. http://www.beckettarchive.org (accessed on 22 October 2015).
- Bryant, J.(2008). Melville Unfolding. Sexuality, Politics, and the Versions of Typee: a Fluid-text Analysis, with an edition of the Typee manuscript. Ann Arbor: University of Michigan Press.
- Elsschot, W. (2007). Achter de Schermen: elektronische editie . Edited by Dirk Van Hulle, Vincent Neyt, and Peter de Bruijn. Kalmthout: Willem Elsschot Genootschap. CD-ROM.
- Ferrer, D. (1998). The open space of the draft page: James Joyce and modern manuscripts. In The Iconic Page in Manuscript, Print, and Digital Culture, edited by George Bornstein and Theresa Tinkle. Ann Arbor: University of Michigan Press, pp: 249–67.
- Melville, H. (2007). Moby-Dick, Edited by John Bryant and Haskell Springer, New York: Pearson / Longman.
- Proust, M. (2012). Autour d’une séquence et des notes du Cahier 46: enjeu du codage dans les brouillons de Proust–Around a Sequence and some Notes of Notebook 46: Encoding Issues about Proust’s Drafts. Edited by Elena Pierazzo and Julie André, with technical support from Raffaele Viglianti. London: King’s College London. http://research.cch.kcl.ac.uk/proust_prototype/ (accessed 9 December 2015).
- Shelley, M. (2013). Frankenstein. In The Shelley-Godwin Archive, directed by Elizabeth C. Denliger and Neil Fraistat. http://www.shelleygodwinarchive.org/ contents/frankenstein (accessed on 25 April 2015).
- Workgroup on Genetic Editions. (2010). An Encoding Model for Genetic Editions. http://www.tei-c.org/Activities/Council/Working/tcw19.html (accessed on 30 October 2015).
- TEI Consortium. (2011). P5 version 2.0 release notes. TEI-C. http://www.tei- c.org/release/doc/tei-p5-doc/readme-2.0.html (accessed on 13 November 2015).
